This whitepaper was developed by contextflow’s Scientific and R&D teams to explain our approach behind lung segmentation. The full text is listed below. For a pdf copy, click here.
Abstract
Automatic lung segmentation in computed tomography (CT) is a critical component of computational medical image analysis. While many approaches exist, it remains a challenging problem for patients whose lungs are affected by disease. Here we describe and evaluate a lung segmentation algorithm that yields high accuracy despite disease patterns thanks to a 3D architecture and diverse training data set. We compare our algorithm with the established state-of-the-art algorithm.
Introduction
Automatic lung segmentation in chest CT scans is relevant for several reasons. It serves as a stepping stone for additional lung-specific analysis such as nodule detection or disease pattern segmentation. Focusing on the lung area helps to improve execution speed of dependent components and to disambiguate any finding in adjacent areas. By itself, lung segmentation yields a measurement of the lung volumetry, which can be meaningful, and is necessary to assess the proportion of disease patterns covering lung tissue.
The existing computer-assisted solutions for lung segmentation make use of a wide range of approaches with different levels of complexity pertaining to various use cases, such as lung cancer screening, COVID-19, or COPD assessment (Hu et al., 2001; Armato et al., 2004; Sluimer et al., 2005; Mansoor et al., 2015; Shamim et al., 2022).
At contextflow, we segment the lungs (chest cavity) by their most inclusive definition: we include pleural cavity patterns such as pneumothorax and effusion in our segmentation, even though they are technically not part of the lungs, and designate them accordingly, since they are relevant for comprehensive lung reporting. Our algorithms segment many pathological patterns if present, including pleural cavity patterns such as pneumothorax and effusion, using this inclusive lung segmentation as a starting point. In practice, lung segmentation needs to be simultaneously fast (because it is only one step in a host of different processes to analyze the lungs) and robust, as our product focuses on the detection and quantification of lung abnormalities, including comparison with former studies, to automatically detect (major) pathological changes.
A 3D approach towards robust lung segmentation in CT
To achieve robust lung segmentation, we first focused on the construction of an adequate database. We selected cases based on their clinical content, trying to cover as many different lung pathologies and anomalies as possible to ensure maximum diversity and extensive pathologic pattern coverage (see annex for details). We also made sure to cover a wide variety of acquisition protocols, since there is a large technical heterogeneity regarding CT vendors and acquisition parameters from different institutions around the world. In these cases, the lungs were manually annotated by expert radiologists. The annotation process consisted of a loop of annotations and quality checks until the annotation passed the quality check.
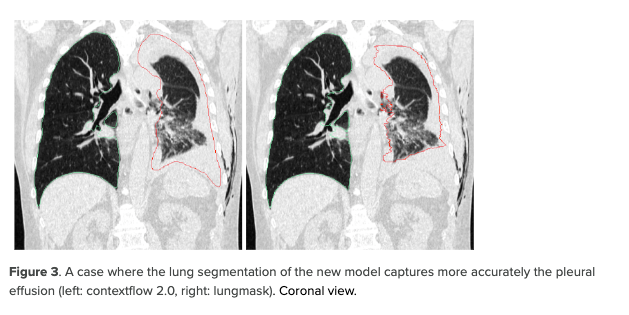
Additionally we worked on the choice of a relevant model architecture. We opted for a 3D-UNet-based architecture. One of the advantages of a model using 3D operations is that 3D information can disambiguate findings hard to discern in a 2D slice. For example, it can be hard to differentiate dense pathological patterns like pleural effusion from structures with soft tissue density like the liver on one slice only. In contrast to 2D models, the 3D model is expected to perform better in these cases, as it uses 3D context to make a prediction. Another advantage is that the model produces a segmentation that is consistent axially by design, contrary to a 2D model that processes each slice independently. It was also designed to be faster, finding a compromise between input size, number of parameters and performance.
Comparative evaluation of lung segmentation accuracy
In order to assess the performance of the developed model, we compared it with a state-of-the-art, publicly-available* model (lungmask) for lung segmentation as a baseline (Hofmanninger et al., 2020). The public model is ranked amongst the best on the public challenge LOLA** (LObe and Lung Analysis) and is frequently used for research purposes for organ segmentation. The model adopts a 2D approach for model training.
We compared the performance between contextflow 2.0 and lungmask on 1722 scans that were sampled from clinical routine without restriction on age, sex, indication or pathology, and that cover findings such as cancer, emphysema, effusion, atelectasis, fibrosis and other pulmonary diseases. For 1694 cases, at least one Region-Of-Interest (ROI) containing a pulmonary pathological pattern is available, while 120 cases have a pixel-wise segmentation mask. All annotations were created by expert radiologists.
Results: segmentation accuracy evaluated by Dice coefficient
We use the widely-adopted Dice coefficient to compare how well the two models’ predictions overlap with an annotation done by a radiologist. The Dice score ranges between 0 (if there is no overlap) and 1 (complete overlap). We report the Dice coefficient for the whole lungs, the right lung and the left lung, measured on the test set in Table 1. We can see that both models achieve very high Dice on average. However, our method tends to achieve higher Dice with a lower standard deviation, which hints to a more robust segmentation performance over the diverse test dataset.
Results: focusing on regions containing disease patterns
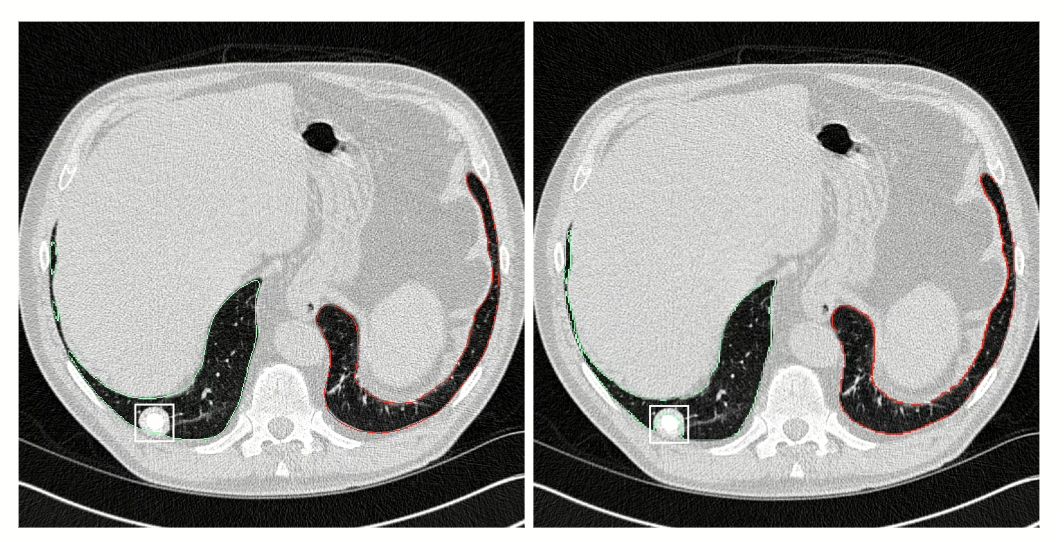
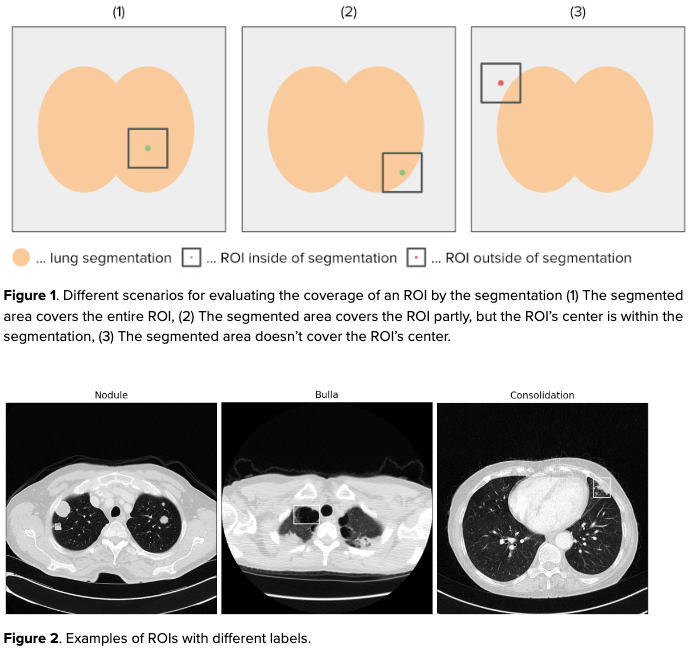
To better assess the difference in performance between the two models, we introduce a metric called Region-Of-Interest (ROI) coverage. ROIs are labeled areas that contain a finding. Here, we consider rectangular ROIs on axial slices containing pulmonary pathological patterns. We define ROI coverage as the percentage of ROI centroids that the lung segmentation is able to cover, meaning that the center of the ROI is included in the segmentation. This metric assesses how accurate the lung segmentation model is, despite the presence of pulmonary diseases. This is crucial in its assessment as a tool to support diagnosis and assessment of lung imaging data.
Annotating ROIs instead of a pixel-wise annotation of pathological patterns is beneficial because it is more time-efficient. In addition, some diffuse lung pathologies are difficult to annotate on a pixel-level even for expert radiologists, so coarsely annotating the entire affected region is more feasible. For this analysis we use ROIs that were created by radiologists on data that cover a wide range of pulmonary pathological patterns.
*https://github.com/JoHof/lungmask **https://lola11.grand-challenge.org/
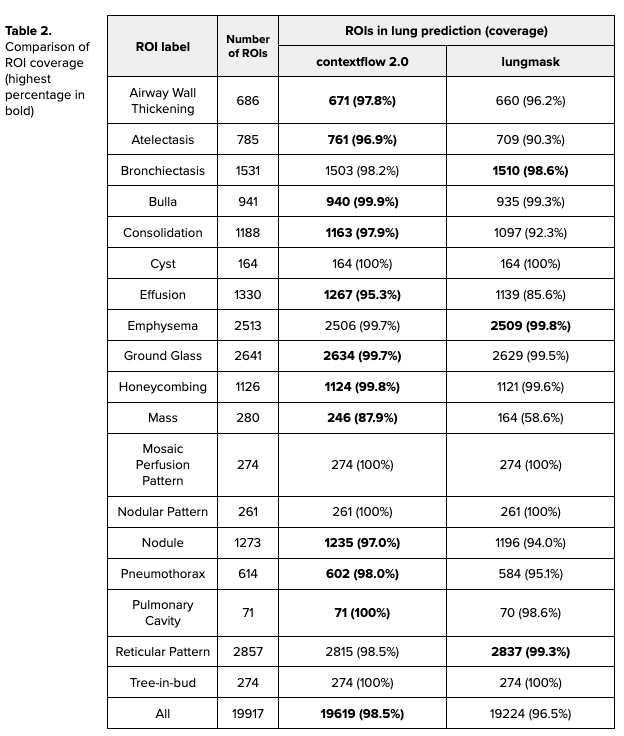
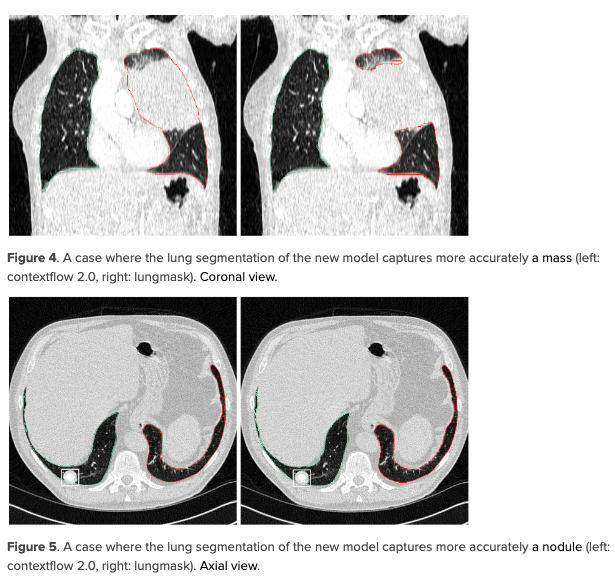
We report the ROI coverage figures in Table 2. The model developed by contextflow performs better overall with a coverage of 98.5% of ROIs compared to 96.5% for the open-source model. This effect is especially pronounced for patterns like masses (87.9% vs. 58.6%) and effusion (95.3% vs. 85.6%). This demonstrates that our model covers the lung pathologies more consistently than the open-source model.
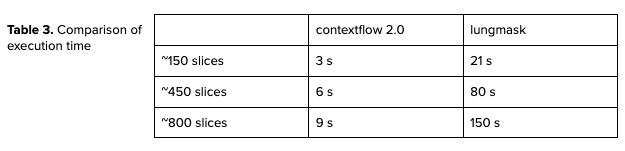
Evaluating execution time
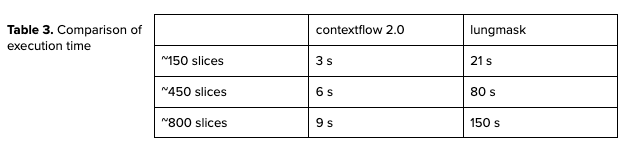
In terms of execution time, our model runs faster than the open-source one. We report the numbers for CPU execution on 8 threads for different sizes of CT scans in Table 3. By design, the open-source algorithm’s execution time directly depends on the number of slices in the input CT whereas the new model depends on the FOV (Field Of View) of the input scan. The developed model can generate lung segmentation much faster than the open-source model.
Summary
In this article we describe how we developed and evaluated a new lung segmentation algorithm. We compared it to an open-source, state-of-the-art solution over which we showed superiority with a conventional metric (Dice coefficient) and with a new clinical-finding-based metric (ROI coverage), showing a higher robustness to a wide variety of cases. Our developed solution also runs faster, making it scalable in terms of the number of scans we can process. Thus, our solution better meets the requirements for usage in clinical practice.
Annex
List of the patterns/pathologies we aim to cover with lung segmentation:
Airway wall thickening, Atelectasis, Bronchiectasis, Bulla, Consolidation, Cyst, Effusion, Emphysema, Ground glass opacification, Honeycombing, Mass, Mosaic attenuation pattern, Nodular pattern, Nodule, Pneumothorax, Pulmonary cavity, Reticular pattern, Tree-in-bud, Fibrosis, Interlobular septal thickening.
Literature
Hofmanninger J, Prayer F, Pan J, Röhrich S, Prosch H, Langs G. Automatic lung segmentation in routine imaging is primarily a data diversity problem, not a methodology problem. European Radiology Experimental. 2020 Dec;4(1):1-3.
Mansoor A, Bagci U, Foster B, Xu Z, Papadakis GZ, Folio LR, Udupa JK, Mollura DJ. Segmentation and image analysis of abnormal lungs at CT: current approaches, challenges, and future trends. Radiographics. 2015 Jul;35(4):1056-76.
Shamim S, Awan MJ, Mohd Zain A, Naseem U, Mohammed MA, Garcia-Zapirain B. Automatic COVID-19 lung infection segmentation through modified unet model. Journal of healthcare engineering. 2022 Apr 11;2022.
Armato III SG, Sensakovic WF. Automated lung segmentation for thoracic CT: impact on computer-aided diagnosis1. Academic Radiology. 2004 Sep 1;11(9):1011-21.
Hu S, Hoffman EA, Reinhardt JM. Automatic lung segmentation for accurate quantitation of volumetric X-ray CT images. IEEE transactions on medical imaging. 2001 Jun;20(6):490-8.
Sluimer I, Prokop M, Van Ginneken B. Toward automated segmentation of the pathological lung in CT. IEEE transactions on medical imaging. 2005 Aug 1;24(8):1025-38.